Ten years ago, I was publishing my first manuscript as 1st author, a manuscript about antimicrobial activity prediction, which would be one of the pillars of Porto Reports. Because of its inaugural character, I choose that article to be subject of the first post on Porto Reports Legacy.
“An SVM model based on physicochemical properties to predict antimicrobial activity from protein sequences with cysteine knot motifs”, this manuscript is far from being a perfect manuscript, but it has some strengths, including an innovative strategy for antimicrobial activity prediction. In fact, the innovation was the main strength for worthing the publication. And by today, I use this work to teach what to do and what not to do in an scientific publication.
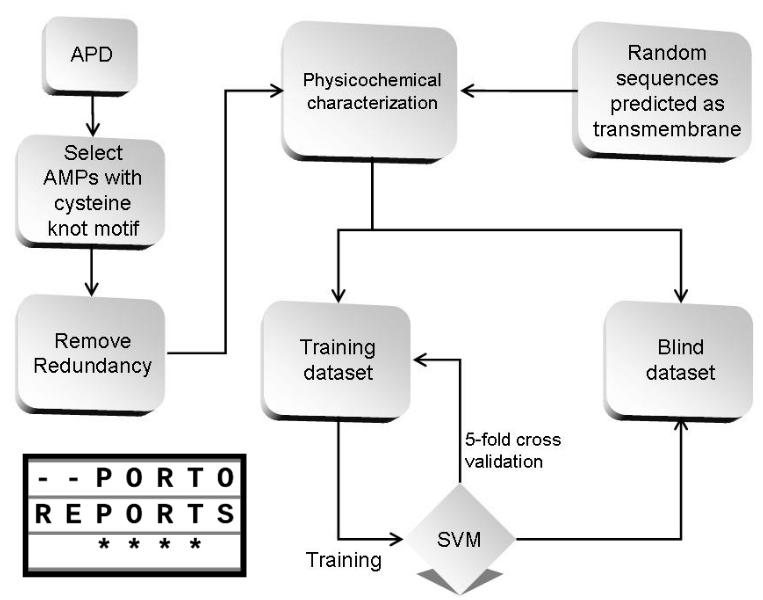
Briefly, the manuscript describes the construction of an antimicrobial activity prediction system using support vector machine as the machine learning algorithm and physicochemical properties as the sequence descriptors. The system reached a good accuracy (~80%) using the polynomial kernel.
The main limitations were (i) the weak English, and in fact, the incorrect use of verb tenses in several manuscript sections; and (ii) the non-contextualization of the computational problem, which makes the manuscript hard to understand for scientists from biological sciences.
Nevertheless, I need to talk about the strengths! To create something innovative, we need to be creative and in 2010 there was a wide field to explore in this topic. In fact, there was only one manuscript until then related to prediction of antimicrobial activity. Thus in this condition, the idea does not need to be bright, it just need to be different.
The difference was not in the algorithm itself, but on how to train the machine learning algorithm. Because the first manuscript demonstrated that there are only slight differences in the predictive power of different algorithms with the same training schemes. Thus, we used physicochemical properties to train the system, reaching a good accuracy. However, there were some limitations on our technique, that were well described and properly addressed, which is always a strength.
This system was the precursor of CS-AMPPred, and due to some errors in the choice of methods, including the support vector machine engine, the original system did not reach its actual potential.
Quality assessment:
Originality: ☆☆☆☆✭
Rigor: ☆☆☆✭✭
Significance to the field: ☆☆☆☆✭
Interest to general audience: ☆☆☆✭✭
Quality of writing: ☆☆✭✭✭
Overall quality of the study: ☆☆☆✭✭
Reference:
Porto W.F., Fernandes F.C., Franco O.L. (2010) An SVM Model Based on Physicochemical Properties to Predict Antimicrobial Activity from Protein Sequences with Cysteine Knot Motifs. In: Ferreira C.E., Miyano S., Stadler P.F. (eds) Advances in Bioinformatics and Computational Biology. BSB 2010. Lecture Notes in Computer Science, vol 6268. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-15060-9_6